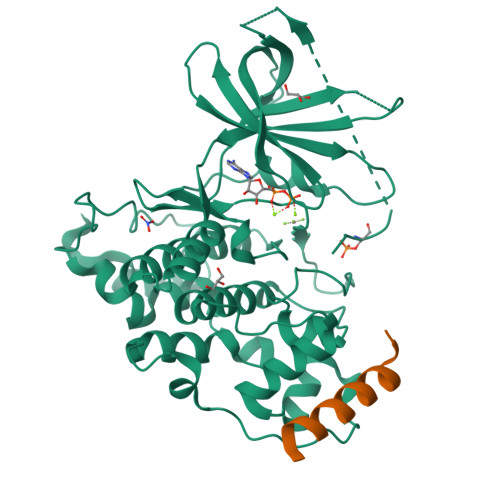
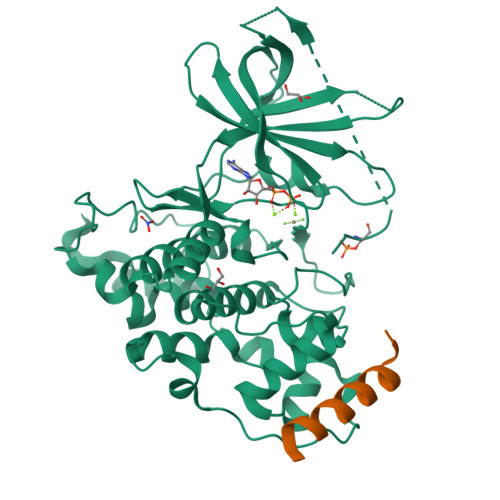
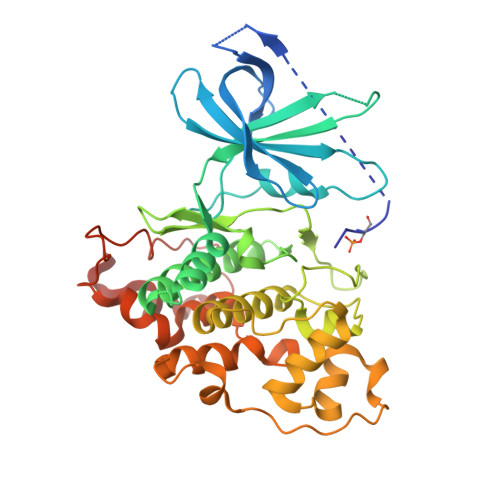
Structural basis of GSK-3 inhibition by N-terminal phosphorylation and by the Wnt receptor LRP6.
Stamos, J.L., Chu, M.L., Enos, M.D., Shah, N., Weis, W.I.(2014) Elife 3: e01998-e01998
- PubMed: 24642411
- DOI: https://doi.org/10.7554/eLife.01998
- Primary Citation of Related Structures:
4NM0, 4NM3, 4NM5, 4NM7, 4NU1 - PubMed Abstract:
Glycogen synthase kinase-3 (GSK-3) is a key regulator of many cellular signaling pathways. Unlike most kinases, GSK-3 is controlled by inhibition rather than by specific activation. In the insulin and several other signaling pathways, phosphorylation of a serine present in a conserved sequence near the amino terminus of GSK-3 generates an auto-inhibitory peptide. In contrast, Wnt/β-catenin signal transduction requires phosphorylation of Ser/Pro rich sequences present in the Wnt co-receptors LRP5/6, and these motifs inhibit GSK-3 activity. We present crystal structures of GSK-3 bound to its phosphorylated N-terminus and to two of the phosphorylated LRP6 motifs. A conserved loop unique to GSK-3 undergoes a dramatic conformational change that clamps the bound pseudo-substrate peptides, and reveals the mechanism of primed substrate recognition. The structures rationalize target sequence preferences and suggest avenues for the design of inhibitors selective for a subset of pathways regulated by GSK-3. DOI: http://dx.doi.org/10.7554/eLife.01998.001.
Organizational Affiliation:
Department of Structural Biology, Stanford University, Stanford, United States.