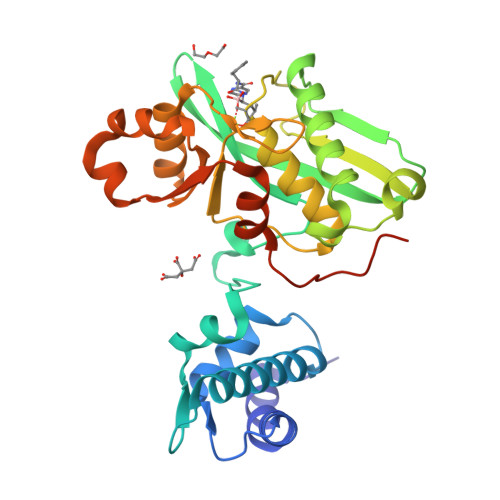
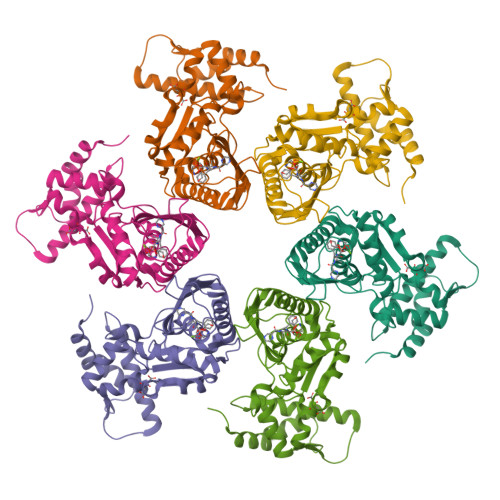
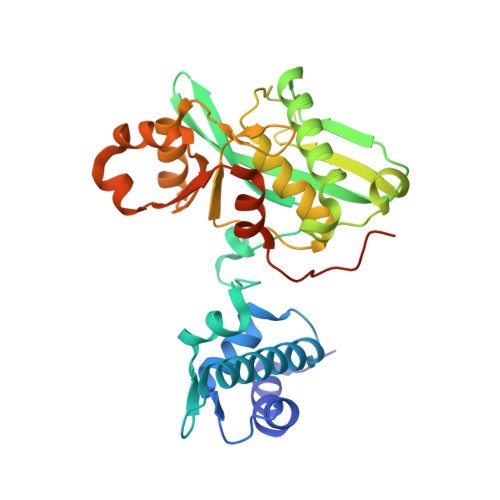
Structural Basis for the Magnesium-Dependent Activation and Hexamerization of the Lon AAA+ Protease
Su, S.-C., Lin, C.-C., Tai, H.-C., Chang, M.-Y., Ho, M.-R., Babu, C.S., Liao, J.-H., Wu, S.-H., Chang, Y.-C., Lim, C., Chang, C.-I.(2016) Structure 24: 676-686
- PubMed: 27041593
- DOI: https://doi.org/10.1016/j.str.2016.03.003
- Primary Citation of Related Structures:
4YPM, 5E7S - PubMed Abstract:
The Lon AAA+ protease (LonA) plays important roles in protein homeostasis and regulation of diverse biological processes. LonA behaves as a homomeric hexamer in the presence of magnesium (Mg(2+)) and performs ATP-dependent proteolysis. However, it is also found that LonA can carry out Mg(2+)-dependent degradation of unfolded protein substrate in an ATP-independent manner. Here we show that in the presence of Mg(2+) LonA forms a non-secluded hexameric barrel with prominent openings, which explains why Mg(2+)-activated LonA can operate as a diffusion-based chambered protease to degrade unstructured protein and peptide substrates efficiently in the absence of ATP. A 1.85 Å crystal structure of Mg(2+)-activated protease domain reveals Mg(2+)-dependent remodeling of a substrate-binding loop and a potential metal-binding site near the Ser-Lys catalytic dyad, supported by biophysical binding assays and molecular dynamics simulations. Together, these findings reveal the specific roles of Mg(2+) in the molecular assembly and activation of LonA.
Organizational Affiliation:
Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan 11529, ROC; Institute of Biochemical Sciences, National Taiwan University, Taipei, Taiwan 10617, ROC.