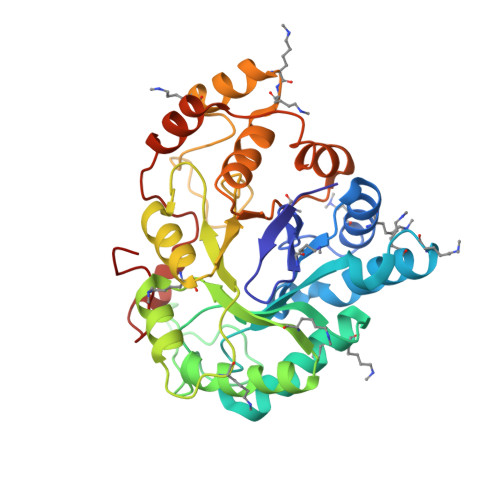
IDD388 Polyhalogenated Derivatives as Probes for an Improved Structure-Based Selectivity of AKR1B10 Inhibitors.
Cousido-Siah, A., Ruiz, F.X., Fanfrlik, J., Gimenez-Dejoz, J., Mitschler, A., Kamlar, M., Vesely, J., Ajani, H., Pares, X., Farres, J., Hobza, P., Podjarny, A.D.(2016) ACS Chem Biol 11: 2693-2705
- PubMed: 27359042
- DOI: https://doi.org/10.1021/acschembio.6b00382
- Primary Citation of Related Structures:
5LIK, 5LIU, 5LIW, 5LIX, 5LIY - PubMed Abstract:
Human enzyme aldo-keto reductase family member 1B10 (AKR1B10) has evolved as a tumor marker and promising antineoplastic target. It shares high structural similarity with the diabetes target enzyme aldose reductase (AR). Starting from the potent AR inhibitor IDD388, we have synthesized a series of derivatives bearing the same halophenoxyacetic acid moiety with an increasing number of bromine (Br) atoms on its aryl moiety. Next, by means of IC 50 measurements, X-ray crystallography, WaterMap analysis, and advanced binding free energy calculations with a quantum-mechanical (QM) approach, we have studied their structure-activity relationship (SAR) against both enzymes. The introduction of Br substituents decreases AR inhibition potency but improves it in the case of AKR1B10. Indeed, the Br atoms in ortho position may impede these drugs to fit into the AR prototypical specificity pocket. For AKR1B10, the smaller aryl moieties of MK181 and IDD388 can bind into the external loop A subpocket. Instead, the bulkier MK184, MK319, and MK204 open an inner specificity pocket in AKR1B10 characterized by a π-π stacking interaction of their aryl moieties and Trp112 side chain in the native conformation (not possible in AR). Among the three compounds, only MK204 can make a strong halogen bond with the protein (-4.4 kcal/mol, using QM calculations), while presenting the lowest desolvation cost among all the series, translated into the most selective and inhibitory potency AKR1B10 (IC 50 = 80 nM). Overall, SAR of these IDD388 polyhalogenated derivatives have unveiled several distinctive AKR1B10 features (shape, flexibility, hydration) that can be exploited to design novel types of AKR1B10 selective drugs.
Organizational Affiliation:
Department of Integrated Structural Biology, Institut de Génétique et de Biologie Moléculaire et Cellulaire, CNRS, INSERM, UdS , 1 rue Laurent Fries 67404 CEDEX Illkirch, France.