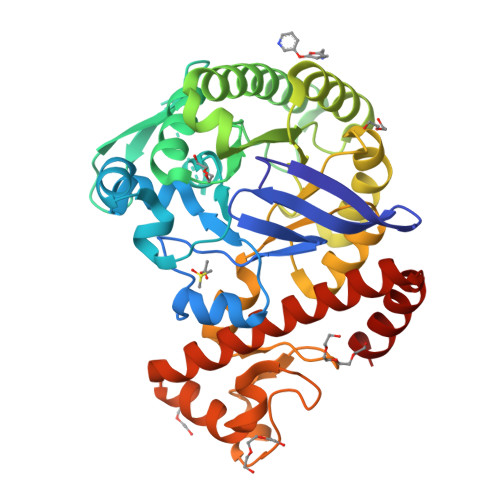
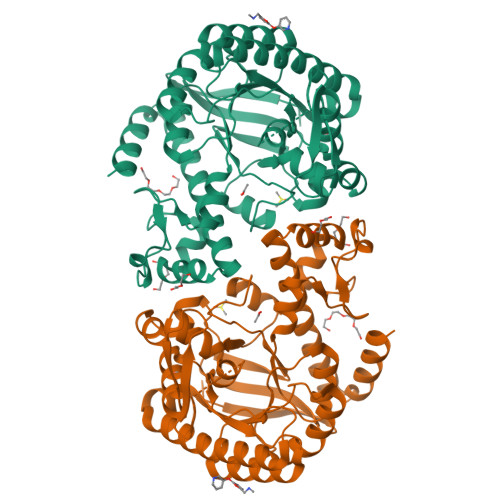
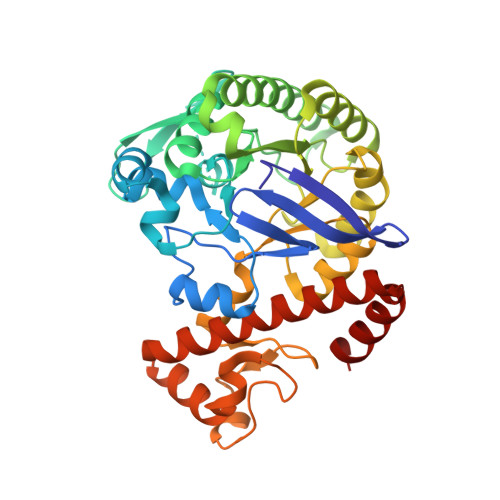
Fragments as Novel Starting Points for tRNA-Guanine Transglycosylase Inhibitors Found by Alternative Screening Strategies.
Hassaan, E., Eriksson, P.O., Geschwindner, S., Heine, A., Klebe, G.(2020) ChemMedChem 15: 324-337
- PubMed: 31808981
- DOI: https://doi.org/10.1002/cmdc.201900604
- Primary Citation of Related Structures:
5N6F, 5SW3, 5UTI, 5UTJ, 5V3C, 6FSO - PubMed Abstract:
Crystallography provides structural information crucial for fragment optimization, however several criteria must be met to screen directly on protein crystals as soakable, well-diffracting specimen must be available. We screened a 96-fragment library against the tRNA-modifying enzyme TGT using crystallography. Eight hits, some with surprising binding poses, were detected. However, the amount of data collection, reduction and refinement is assumed substantial. Therefore, having a reliable cascade of fast and cost-efficient methods available for pre-screening before embarking to elaborate crystallographic screening appears beneficial. This allows filtering of compounds to the most promising hits, available to rapidly progress from hit-to-lead. But how to ensure that this workflow is reliable? To answer this question, we also applied SPR and NMR to the same screening sample to study whether identical hits are retrieved. Upon hit-list comparisons, crystallography shows with NMR and SPR, only one overlapping hit and all three methods shared no common hits. This questions a cascade-type screening protocol at least in the current example. Compared to crystallography, SPR and NMR detected higher percentages of non-active-site binders suggesting the importance of running reporter ligand-based competitive screens in SPR and NMR, a requirement not needed in crystallography. Although not specific, NMR proved a more sensitive method relative to SPR and crystallography, as it picked up the highest numbers of binders.
Organizational Affiliation:
Institute of Pharmaceutical Chemistry, Philipps University Marburg, Marbacher Weg 6, 35032, Marburg, Germany.