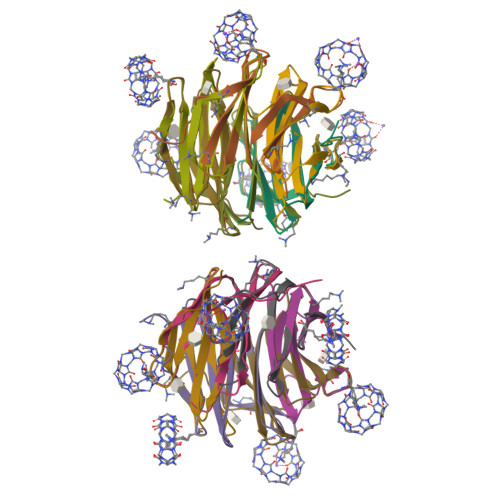
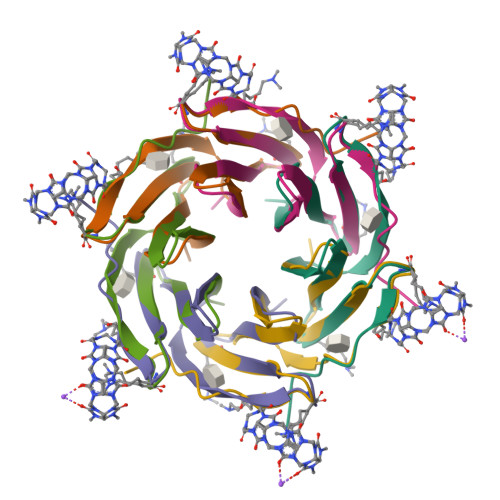
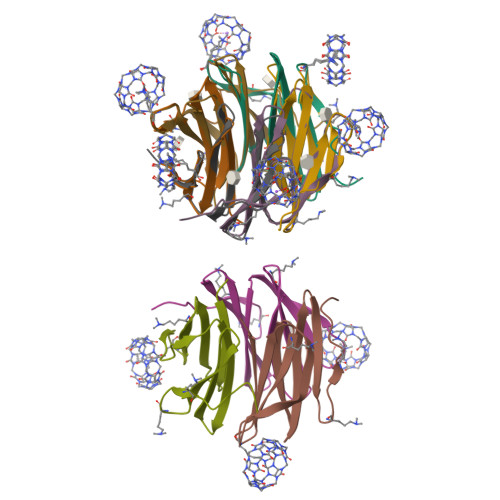
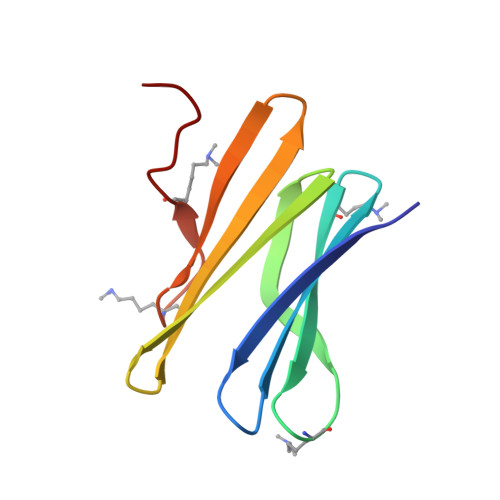
Engineered assembly of a protein-cucurbituril biohybrid.
Guagnini, F., Engilberge, S., Ramberg, K.O., Perez, J., Crowley, P.B.(2020) Chem Commun (Camb) 56: 360-363
- PubMed: 31825399
- DOI: https://doi.org/10.1039/c9cc07198a
- Primary Citation of Related Structures:
6STZ, 6SU0 - PubMed Abstract:
A crystalline biohybrid with a 4 : 1 protein : cucurbituril mass ratio is presented. This result was achieved by engineering additional cucurbit[7]uril (Q7) binding sites into a β-propeller protein. In contrast to the parent protein, Q7-controlled assembly of the engineered variant occurred in solution, as evidenced by NMR and SAXS measurements.
Organizational Affiliation:
School of Chemistry, National University of Ireland Galway, University Road, Galway, H91 TK33, Ireland. peter.crowley@nuigalway.ie.