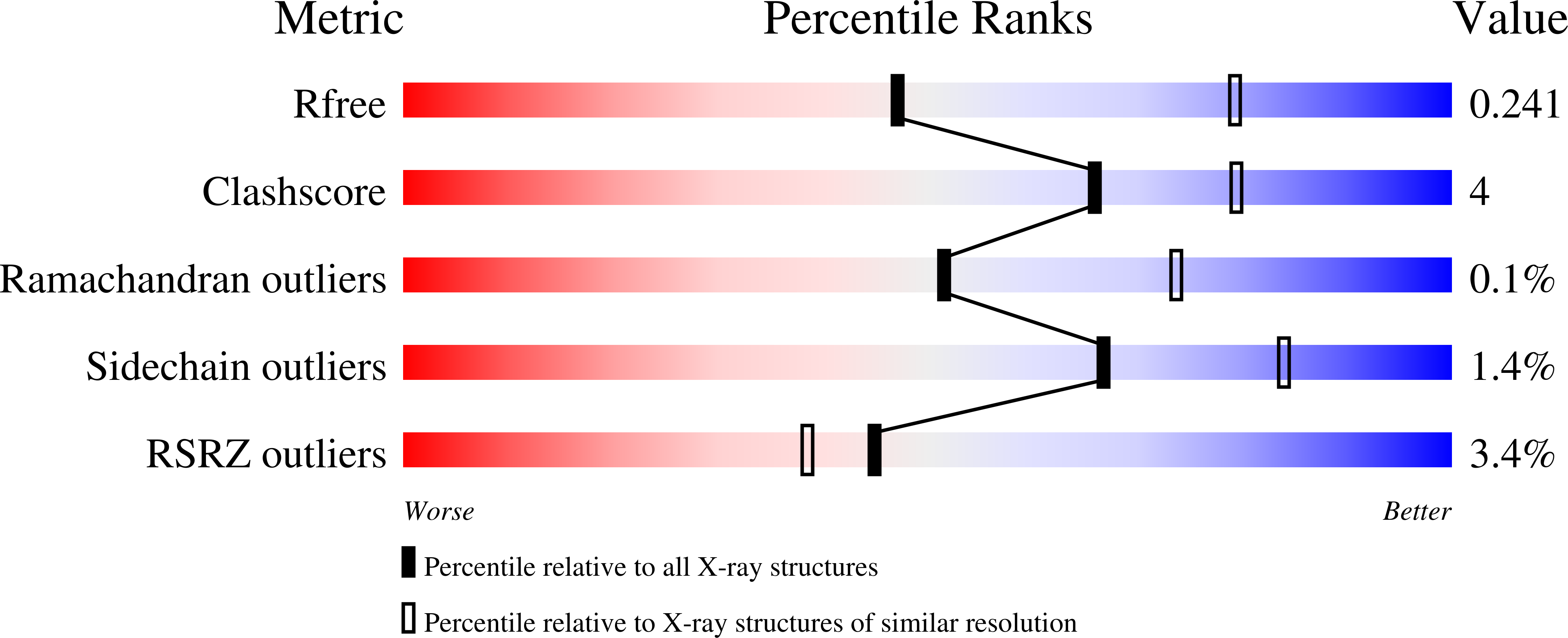
The antigenic anatomy of SARS-CoV-2 receptor binding domain.
Dejnirattisai, W., Zhou, D., Ginn, H.M., Duyvesteyn, H.M.E., Supasa, P., Case, J.B., Zhao, Y., Walter, T.S., Mentzer, A.J., Liu, C., Wang, B., Paesen, G.C., Slon-Campos, J., Lopez-Camacho, C., Kafai, N.M., Bailey, A.L., Chen, R.E., Ying, B., Thompson, C., Bolton, J., Fyfe, A., Gupta, S., Tan, T.K., Gilbert-Jaramillo, J., James, W., Knight, M., Carroll, M.W., Skelly, D., Dold, C., Peng, Y., Levin, R., Dong, T., Pollard, A.J., Knight, J.C., Klenerman, P., Temperton, N., Hall, D.R., Williams, M.A., Paterson, N.G., Bertram, F.K.R., Siebert, C.A., Clare, D.K., Howe, A., Radecke, J., Song, Y., Townsend, A.R., Huang, K.A., Fry, E.E., Mongkolsapaya, J., Diamond, M.S., Ren, J., Stuart, D.I., Screaton, G.R.(2021) Cell 184: 2183
- PubMed: 33756110
- DOI: https://doi.org/10.1016/j.cell.2021.02.032
- Primary Citation of Related Structures:
7BEH, 7BEI, 7BEJ, 7BEK, 7BEL, 7BEM, 7BEN, 7BEO, 7BEP, 7ND3, 7ND4, 7ND5, 7ND6, 7ND7, 7ND8, 7ND9, 7NDA, 7NDB, 7NDC, 7NDD - PubMed Abstract:
Antibodies are crucial to immune protection against SARS-CoV-2, with some in emergency use as therapeutics. Here, we identify 377 human monoclonal antibodies (mAbs) recognizing the virus spike and focus mainly on 80 that bind the receptor binding domain (RBD). We devise a competition data-driven method to map RBD binding sites. We find that although antibody binding sites are widely dispersed, neutralizing antibody binding is focused, with nearly all highly inhibitory mAbs (IC 50 < 0.1 μg/mL) blocking receptor interaction, except for one that binds a unique epitope in the N-terminal domain. Many of these neutralizing mAbs use public V-genes and are close to germline. We dissect the structural basis of recognition for this large panel of antibodies through X-ray crystallography and cryoelectron microscopy of 19 Fab-antigen structures. We find novel binding modes for some potently inhibitory antibodies and demonstrate that strongly neutralizing mAbs protect, prophylactically or therapeutically, in animal models.
Organizational Affiliation:
Wellcome Centre for Human Genetics, Nuffield Department of Medicine, University of Oxford, Oxford OX3 7BN, UK.