RCSB PDB Help
Search and Browse > Advanced Search
Chemical Sketch Tool
Introduction
What does the Chemical Sketch Tool do?
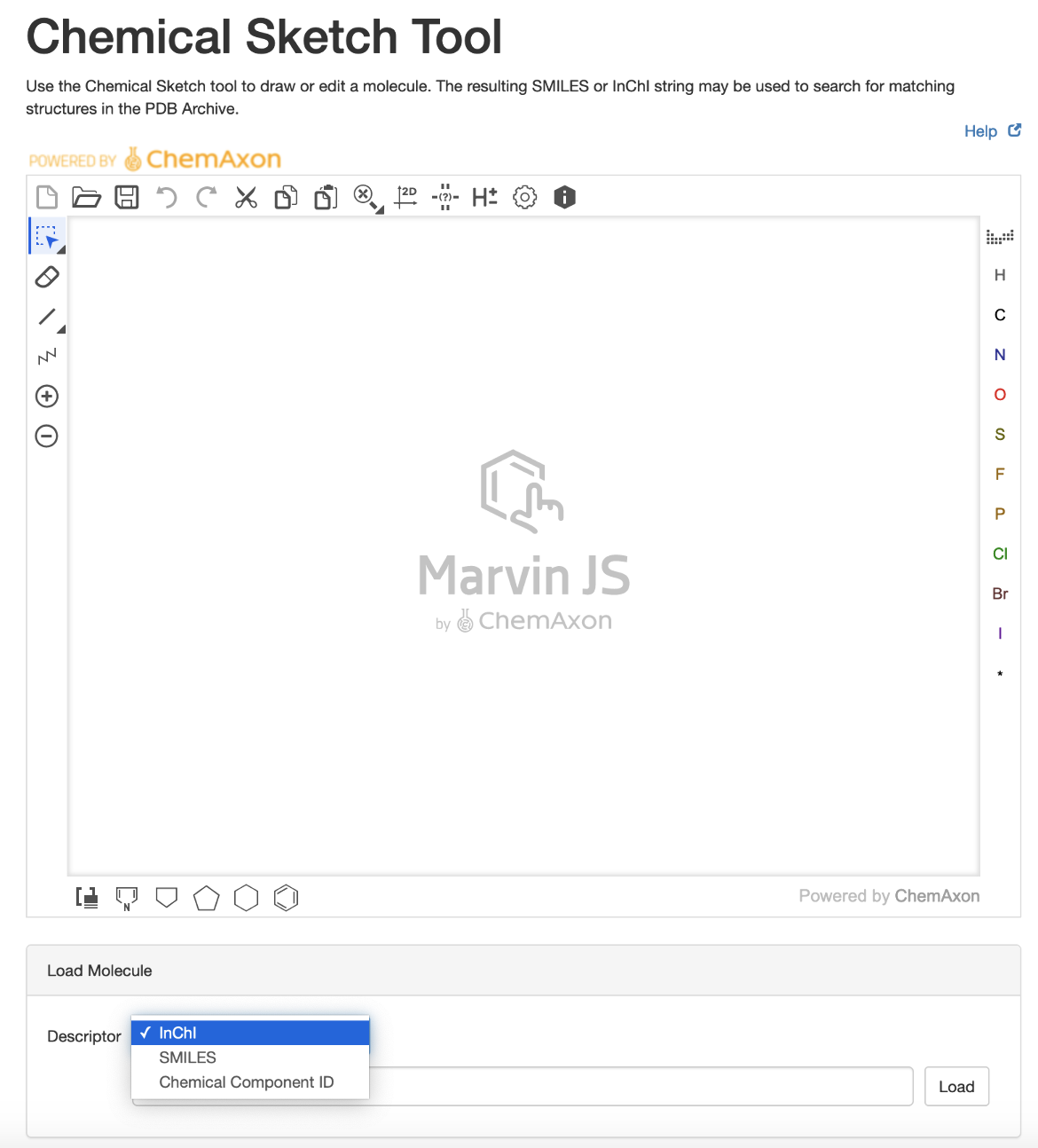
The Chemical Sketch Tool uses Marvin JS, a web-based chemical sketcher developed by ChemAxon, to quickly and conveniently draw 2D chemical structures from scratch, chemical descriptors (SMILES or InChI), or ligand codes in the wwPDB Chemical Component Dictionary (CCD). The tool allows users to edit the 2D chemical drawing as needed to automatically generate new chemical descriptors. The user can then use the descriptor to query the PDB archive for the specific ligand molecule or similar ones.
Why is the Chemical Sketch Tool useful?
You can search the PDB archive for a specific ligand or similar ligands based on the 2D chemical drawing of a molecule. In doing so you can use the graphical interface to draw a molecule of interest or load either the SMILES or InChI descriptors. Both chemical descriptors (SMILES and InChI) of the molecule drawn will be automatically generated and will be available to use as a query. You can also edit a molecule that is drawn or loaded into the tool to add or remove atoms or groups of atoms and then use the new molecule to query the PDB archive.
Documentation
Tool Interface
The Chemical Sketch Tool can be accessed in two ways:
1. By clicking on the "Open Chemical Sketch Tool" button in the Chemical Search options in the Advanced Search.

|
| Access to the Chemical Sketch tool from the Advanced Search Options - click on the button labelled "Open Chemical Sketch Tool" |
2. By directly clicking the Chemical Sketch Tool link here
How to use the Chemical Sketch Tool?
Draw the 2D chemical structure
There are several ways to make a 2D chemical drawing of a molecule:
- Draw the molecule of interest using the options available in the Marvin JS tool
- A built-in menu of chemical groups, chains, and rings, along with several keyboard shortcuts provide quick access to the most common editing features for drawing even large and complex molecules.
- This option is useful for novel ligands that are not present in the CCD.
- Detailed help documentation for using the Marvin JS graphic editor is available can be found here:
- Load the molecule of interest (or a closely related one) using chemical descriptors (e.g., InChI or SMILES) or the Chemical Component ID (e.g., ATP, TTT)
- This is useful when you have access to chemical descriptors of similar ligands or their substructure elements
Edit the 2D Chemical Drawing
Once the 2D structure of the molecule is drawn or loaded, you can add/remove atoms or groups of atoms using the Marvin JS graphic editor.
As you edit the molecule, its chemical descriptors (InChI and SMILES) are automatically updated and a "Search" panel appears at the bottom of the page.
Chemical Search
- Select the relevant search type from the “Search the PDB” pulldown options. Learn more about the Match types - "Similar Ligands (Stereospecific)", "Substructure (Stereospecific)", "Exact match", etc. in the documentation for Chemical Search.
- Select InChI or SMILES radio buttons so that the selected descriptor is passed to the Advanced Search as an input along with the selected Match Type parameter.
- The search results pages show all CCD and BIRD molecules that match the query parameters.
- If you want to refine your search (i.e., edit your chemical drawing or select other match type options), scroll up to the Advanced Search query details sections and click on the “Open Chemical Sketch Tool” button. This will return you to the Chemical Sketch Tool. You can now make further edits to the 2D chemical drawing, select a different match type, and/or select a different chemical descriptor to re-run your search.
- Note: For some molecules the chemical search results returned from the standard InChI may provide greater specificity compared to a search initiated using a SMILES chemical descriptor. Depending on whether you wish to see fewer or more matches in your results you can use the appropriate descriptor for the search.
Examples
1. Search for a molecular definition that exactly matches an InChI
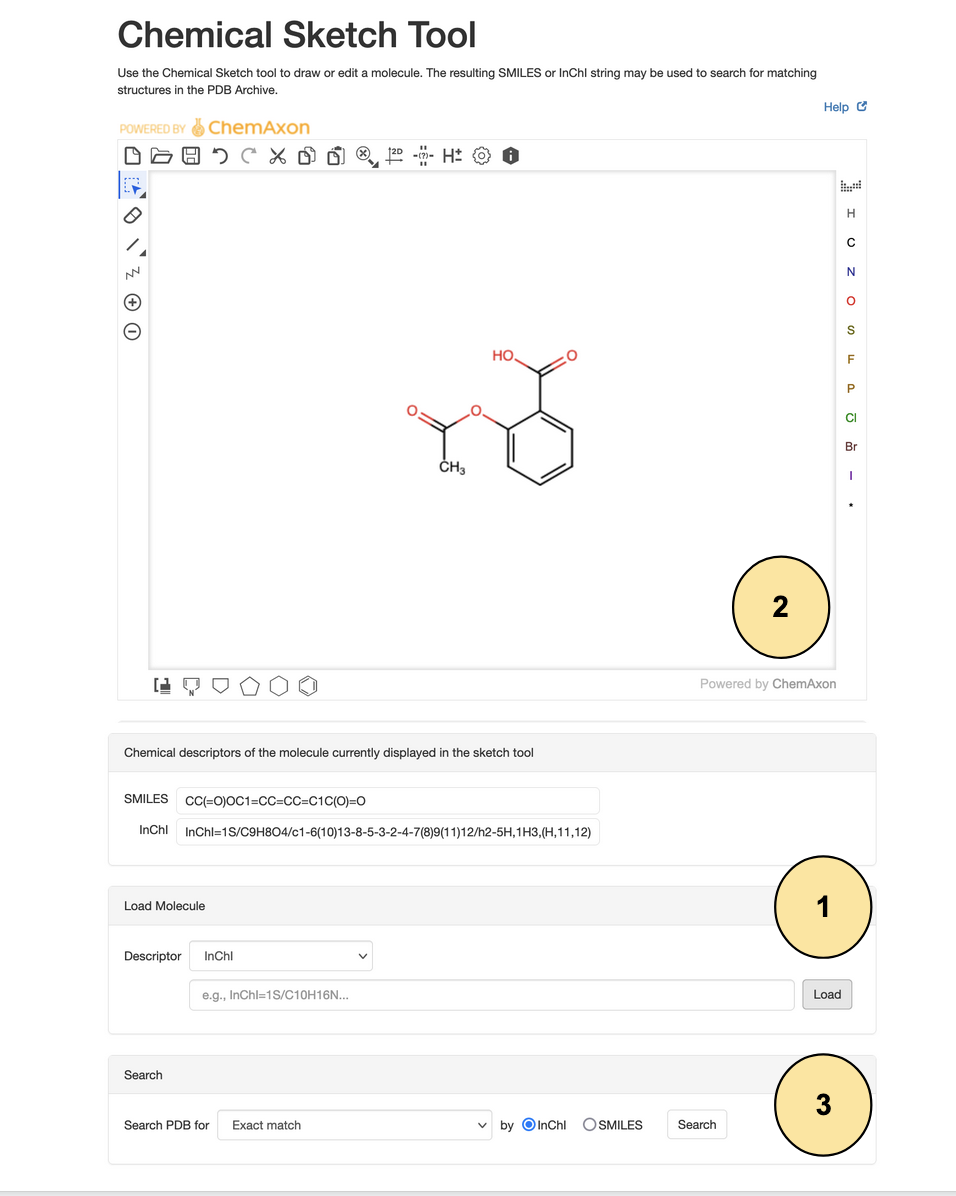
- Say you found a chemical descriptor, such as InChI=1S/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12), in a textbook, scientific literature, or a cheminformatics resource. Copy the entire descriptor and paste it in the "Load Molecule" panel of the Chemical Sketch Tool. Click Load
- A 2D drawing of the molecule is displayed by the sketch tool. Now you know what the molecule looks like.
- To find if this molecule is present in the wwPDB CCD, scroll down and in the Search panel select the search options
- for match type choose "Exact Match"
- for chemical descriptor choose InChI
Click on the Search button to get Search Results
2. Search for molecular definitions that are similar to the CCD ID TTT
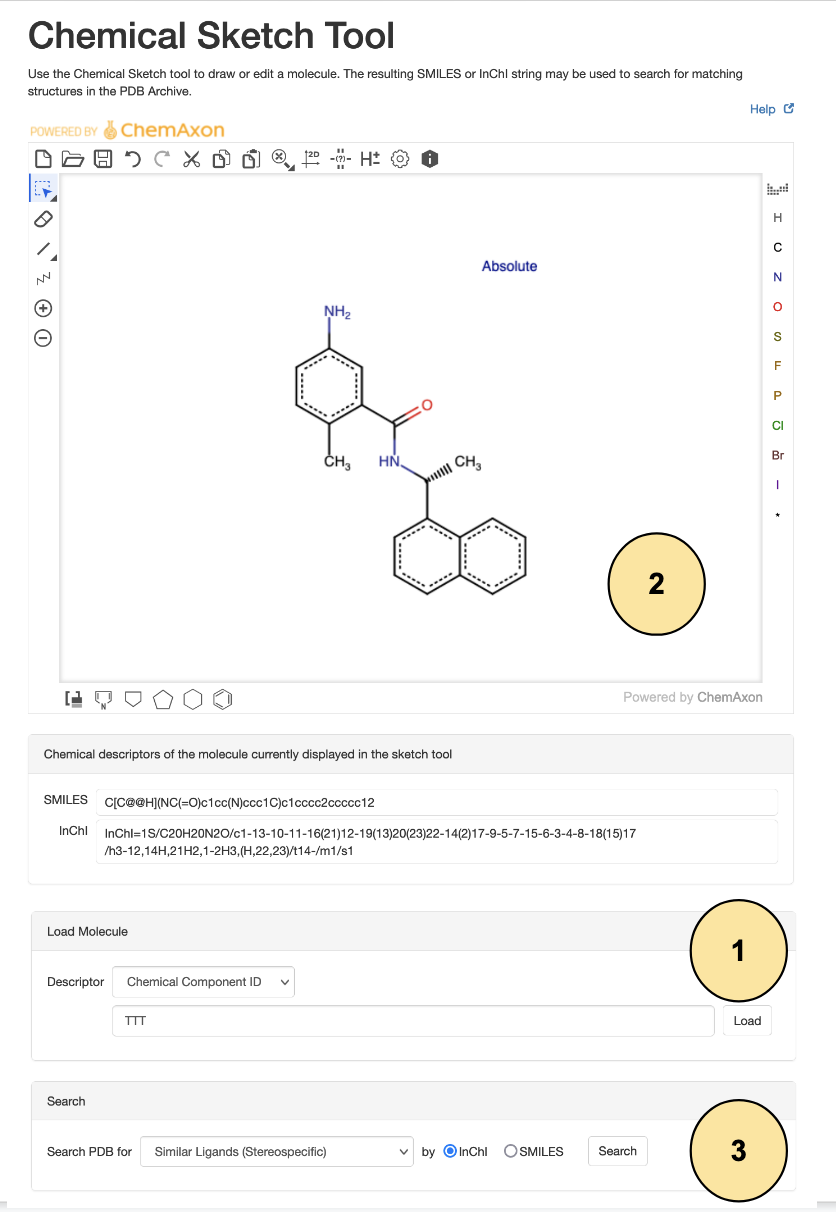
- Open the Chemical Sketch Tool and in the "Load Molecule" panel, select "Chemical Component ID" and enter "TTT", then click on the "Load" button.
- Scroll down to the bottom of the page and select the search options
- for match type choose Similar Ligands (Stereospecific)
- for chemical descriptor choose InChI
- Click on Search and get the Search Results
3. Search for molecular definitions that are substructures of a modified version of CCD ID FMN
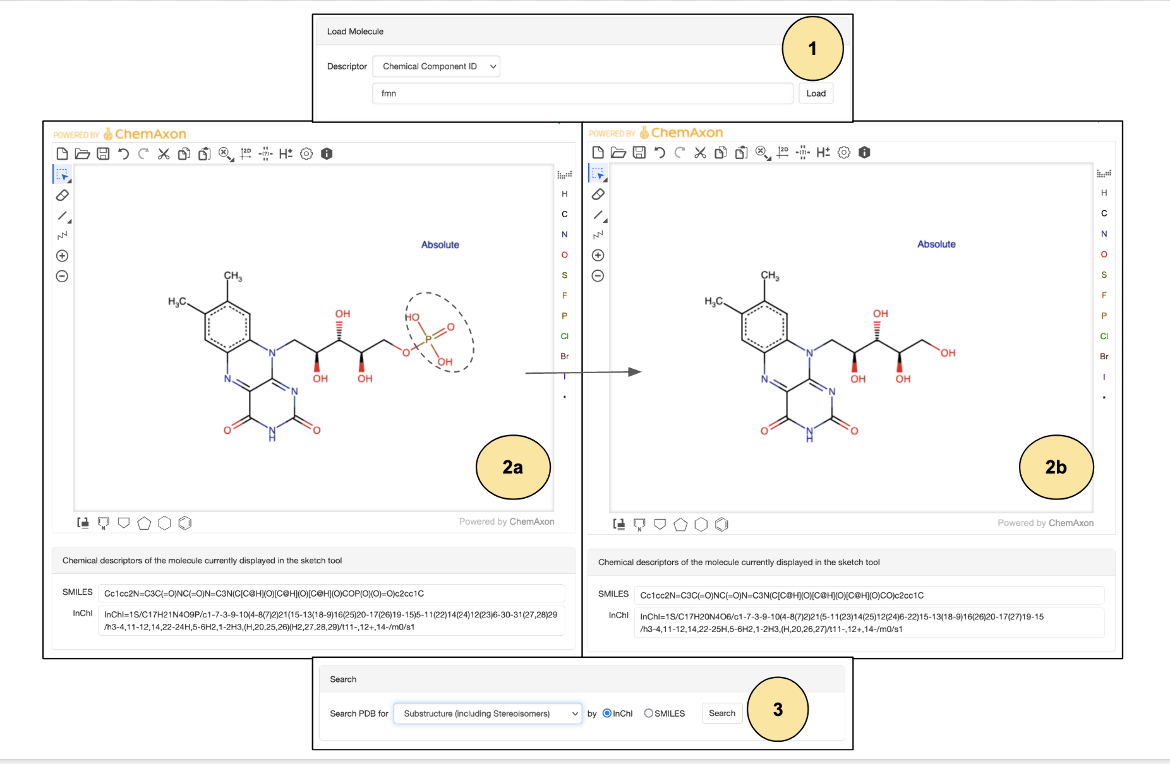
- Open the Chemical Sketch Tool and in the "Load Molecule" panel, select "Chemical Component ID" and enter "FMN", then click on the "Load" button.
- Edit and remove the entire phosphate group for the search. Note that Chemical Descriptors are updated.
- After selecting "Substructure (including Stereoisomers)" click on the Search button to run a search for the modified molecule and get Search Results