News
Structure Predictors: Use ModelCIF for Computed Structure Models
01/30 
ModelCIF (GitHub) is a data information framework developed for and by computational structural biologists to describe structural models of macromolecules derived from computational methods. It provides an extensible data representation for deposition, archiving, and public dissemination of these models of proteins to enable delivery of Findable, Accessible, Interoperable, and Reusable (FAIR) data to users worldwide.
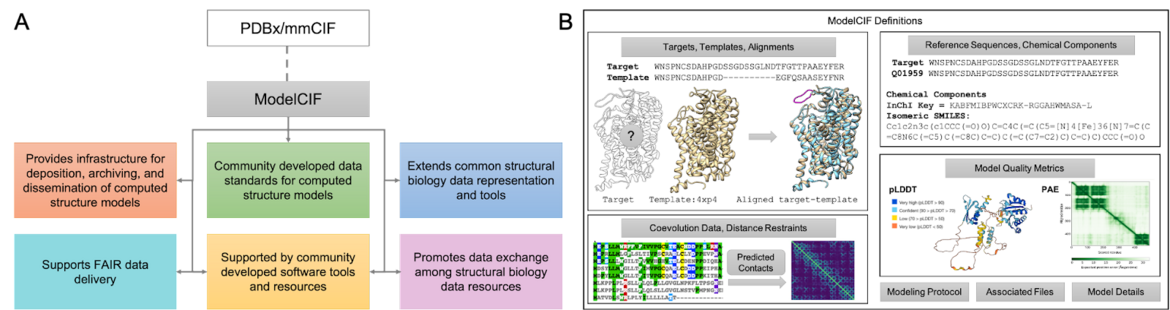 A. Overview of the ModelCIF extension of PDBx/mmCIF. B. Schematic representation of ModelCIF data specifications. ModelCIF includes definitions for input data used in template-based and template-free modeling; reference information for macromolecular sequences and small molecule components; local and global CSM quality metrics; and metadata information regarding modeling protocol, CSM classification (ab initio, homology, etc.) and descriptions of associated files.
A. Overview of the ModelCIF extension of PDBx/mmCIF. B. Schematic representation of ModelCIF data specifications. ModelCIF includes definitions for input data used in template-based and template-free modeling; reference information for macromolecular sequences and small molecule components; local and global CSM quality metrics; and metadata information regarding modeling protocol, CSM classification (ab initio, homology, etc.) and descriptions of associated files.ModelCIF is an extension of the Protein Data Bank Exchange/macromolecular Crystallographic Information Framework (PDBx/mmCIF), which is the global data standard for representing experimentally-determined, three-dimensional (3D) structures of macromolecules and associated metadata. The PDBx/mmCIF framework and its extensions (e.g., ModelCIF) are managed by the wwPDB in collaboration with relevant community stakeholders such as the wwPDB ModelCIF Working Group.
This semantically rich and extensible data framework for representing computed structure models (CSMs) accelerates the pace of scientific discovery. Furthermore, use of this data standard promotes interoperation among structural biology data resources, with ModelCIF currently used by the ModelArchive, AlphaFold DB, and MODBASE repositories. A manuscript was recently submitted to bioRxiv describing the architecture, contents, and governance of ModelCIF as well as tools and processes for maintaining and extending the data standard [1].
Visit the ModelCIF GitHub for more information about this data information framework.
[1} Vallat B, Tauriello G, Bienert S, Haas J, Webb BM, et al. ModelCIF: An extension of PDBx/mmCIF data representation for computed structure models. bioRxiv doi: 10.1101/2022.12.06.518550.












