The essential function of ISCU2 and its conserved N-terminus in Fe/S cluster biogenesis
Freibert, S.A., Boniecki, M.T., Shulz, V., Wilbrecht, C., Krapoth, N., Muhlenhoff, U., Stehling, O., Cygler, M., Lill, R.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cysteine desulfurase, mitochondrial | 406 | Homo sapiens | Mutation(s): 0 Gene Names: NFS1, NIFS, HUSSY-08 EC: 2.8.1.7 | 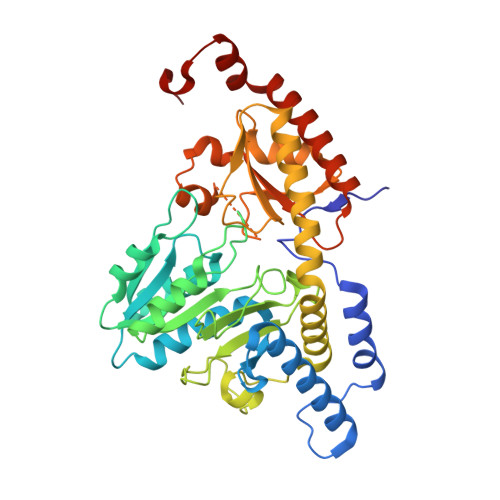 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9Y697 (Homo sapiens) Explore Q9Y697 Go to UniProtKB: Q9Y697 | |||||
PHAROS: Q9Y697 GTEx: ENSG00000244005 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Y697 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| LYR motif-containing protein 4 | 91 | Homo sapiens | Mutation(s): 1 Gene Names: LYRM4, C6orf149, ISD11, CGI-203 | 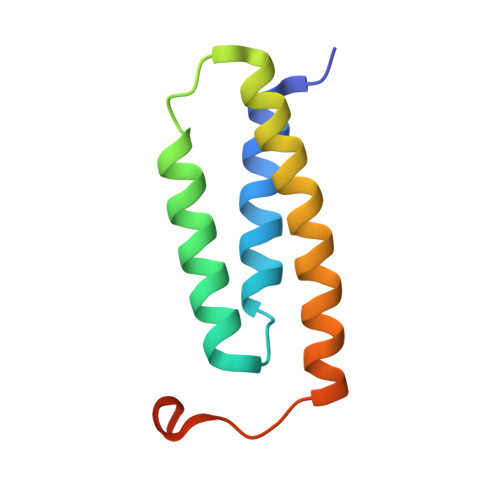 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9HD34 (Homo sapiens) Explore Q9HD34 Go to UniProtKB: Q9HD34 | |||||
PHAROS: Q9HD34 GTEx: ENSG00000214113 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9HD34 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Acyl carrier protein | 77 | Escherichia coli | Mutation(s): 0 | 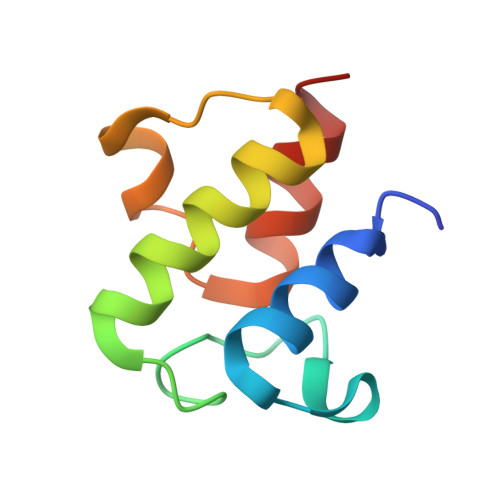 | |
UniProt | |||||
Find proteins for P0A6A8 (Escherichia coli (strain K12)) Explore P0A6A8 Go to UniProtKB: P0A6A8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A6A8 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Iron-sulfur cluster assembly enzyme ISCU, mitochondrial | 143 | Homo sapiens | Mutation(s): 0 Gene Names: ISCU, NIFUN | 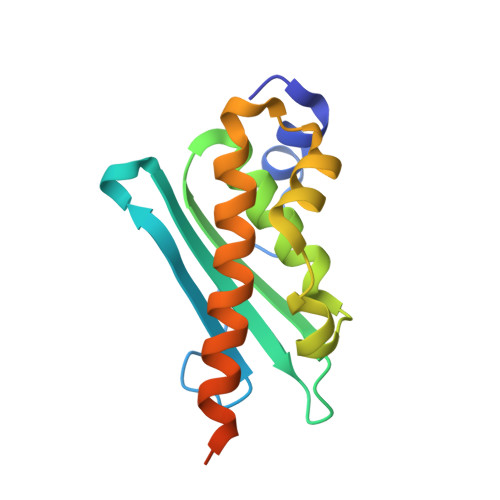 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9H1K1 (Homo sapiens) Explore Q9H1K1 Go to UniProtKB: Q9H1K1 | |||||
PHAROS: Q9H1K1 GTEx: ENSG00000136003 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9H1K1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 12 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 8Q1 Query on 8Q1 | RB [auth C] | S-[2-({N-[(2R)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-beta-alanyl}amino)ethyl] dodecanethioate C23 H45 N2 O8 P S MVHUOSAYFQKAMT-NRFANRHFSA-N |  | ||
| P15 Query on P15 | DA [auth A] | 2,5,8,11,14,17-HEXAOXANONADECAN-19-OL C13 H28 O7 FHHGCKHKTAJLOM-UHFFFAOYSA-N |  | ||
| EDT Query on EDT | NB [auth B] | {[-(BIS-CARBOXYMETHYL-AMINO)-ETHYL]-CARBOXYMETHYL-AMINO}-ACETIC ACID C10 H16 N2 O8 KCXVZYZYPLLWCC-UHFFFAOYSA-N |  | ||
| PLP Query on PLP | E [auth A] | PYRIDOXAL-5'-PHOSPHATE C8 H10 N O6 P NGVDGCNFYWLIFO-UHFFFAOYSA-N |  | ||
| 1PE Query on 1PE | YB [auth D] | PENTAETHYLENE GLYCOL C10 H22 O6 JLFNLZLINWHATN-UHFFFAOYSA-N |  | ||
| MES Query on MES | SB [auth C] | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID C6 H13 N O4 S SXGZJKUKBWWHRA-UHFFFAOYSA-N |  | ||
| PG4 Query on PG4 | W [auth A], X [auth A] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| DTT Query on DTT | KA [auth A] | 2,3-DIHYDROXY-1,4-DITHIOBUTANE C4 H10 O2 S2 VHJLVAABSRFDPM-IMJSIDKUSA-N |  | ||
| PGE Query on PGE | LA [auth A], UA [auth A], VA [auth A] | TRIETHYLENE GLYCOL C6 H14 O4 ZIBGPFATKBEMQZ-UHFFFAOYSA-N |  | ||
| PEG Query on PEG | AB [auth A] BB [auth A] EA [auth A] IA [auth A] JA [auth A] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | AC [auth D] CB [auth A] F [auth A] G [auth A] H [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | AA [auth A] BA [auth A] BC [auth D] CA [auth A] CC [auth D] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 86.348 | α = 90 |
| b = 86.348 | β = 90 |
| c = 245.359 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| PHASER | phasing |